Abstract
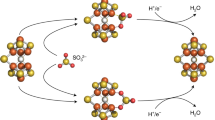
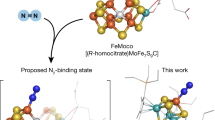
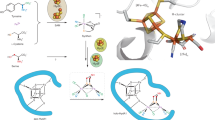
Molybdenum nitrogenase catalyses the ambient reduction of N2 to NH3 at the M-cluster, a complex cofactor that comprises two metal-sulfur partial cubanes ligated by an interstitial carbide and three belt-sulfurs. A recent crystallographic study suggests binding of N2 via displacement of the belt-sulfur(s) of the M-cluster upon turnover. However, direct proof of N2 binding and belt-sulfur mobilization during catalysis remains elusive. Here we show that N2 is captured on the M-cluster via electron and sulfur depletion, and that the N2-captured state is catalytically competent in generating NH3. Moreover, we demonstrate that product release occurs only when sulfite is supplied along with a reductant, that sulfite is inserted as sulfide into the belt-sulfur-displaced positions and that there is a dynamic in-and-out of belt-sulfurs during catalysis. Together, these results establish the mobilization of cofactor belt-sulfurs as a crucial, yet overlooked, mechanistic element of the nitrogenase reaction.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings of this study are available within the article and its Supplementary Information files, or from the corresponding authors upon reasonable request. The structural data related to this work are available at the Worldwide Protein Data Bank (http://www.wwpdb.org) with PDB accession codes 7MCI, 6UG0 and 6VXT. Source data are provided with this paper.
References
Burgess, B. K. & Lowe, D. J. Mechanism of molybdenum nitrogenase. Chem. Rev. 96, 2983–3012 (1996).
Buscagan, T. M. & Rees, D. C. Rethinking the nitrogenase mechanism: activating the active site. Joule 3, 2662–2678 (2019).
Rutledge, H. L. & Tezcan, F. A. Electron transfer in nitrogenase. Chem. Rev. 120, 5158–5193 (2020).
Jasniewski, A. J., Lee, C. C., Ribbe, M. W. & Hu, Y. Reactivity, mechanism, and assembly of the alternative nitrogenases. Chem. Rev. 120, 5107–5157 (2020).
Hu, Y., Lee, C. C. & Ribbe, M. W. Extending the carbon chain: hydrocarbon formation catalysed by vanadium/molybdenum nitrogenases. Science 333, 753–755 (2011).
Lee, C. C., Hu, Y. & Ribbe, M. W. Vanadium nitrogenase reduces CO. Science 329, 642 (2010).
Seefeldt, L. C. et al. Reduction of substrates by nitrogenases. Chem. Rev. 120, 5082–5106 (2020).
Einsle, O. & Rees, D. C. Structural enzymology of nitrogenase enzymes. Chem. Rev. 120, 4969–5004 (2020).
Spatzal, T. et al. Evidence for interstitial carbon in nitrogenase FeMo cofactor. Science 334, 940 (2011).
Wiig, J. A., Hu, Y., Lee, C. C. & Ribbe, M. W. Radical SAM-dependent carbon insertion into the nitrogenase M-cluster. Science 337, 1672–1675 (2012).
Lowe, D. J. & Thorneley, R. N. F. The mechanism of Klebsiella pneumoniae nitrogenase action. Pre-steady-state kinetics of H2 formation. Biochem. J. 224, 877–886 (1984).
Lowe, D. J. & Thorneley, R. N. F. The mechanism of Klebsiella pneumoniae nitrogenase action. The determination of rate constants required for the simulation of the kinetics of N2 reduction and H2 evolution. Biochem. J. 224, 895–901 (1984).
Thorneley, R. N. F. & Lowe, D. J. The mechanism of Klebsiella pneumoniae nitrogenase action. Pre-steady-state kinetics of an enzyme-bound intermediate in N2 reduction and of NH3 formation. Biochem. J. 224, 887–894 (1984).
Thorneley, R. N. F. & Lowe, D. J. The mechanism of Klebsiella pneumoniae nitrogenase action. Simulation of the dependences of H2-evolution rate on component-protein concentration and ratio and sodium dithionite concentration. Biochem. J. 224, 903–909 (1984).
Lee, S. C., Lo, W. & Holm, R. H. Developments in the biomimetic chemistry of cubane-type and higher nuclearity iron-sulphur clusters. Chem. Rev. 114, 3579–3600 (2014).
Spatzal, T., Perez, K. A., Einsle, O., Howard, J. B. & Rees, D. C. Ligand binding to the FeMo-cofactor: structures of CO-bound and reactivated nitrogenase. Science 345, 1620–1623 (2014).
Spatzal, T., Perez, K. A., Howard, J. B. & Rees, D. C. Catalysis-dependent selenium incorporation and migration in the nitrogenase active site iron-molybdenum cofactor. eLife 4, e11620 (2015).
Kang, W., Lee, C. C., Jasniewski, A. J., Ribbe, M. W. & Hu, Y. Structural evidence for a dynamic metallocofactor during N2 reduction by Mo-nitrogenase. Science 368, 1381–1385 (2020).
Yates, M. G. In Biological Nitrogen Fixation (eds Stacey, G., Burris, R. H. & Evans, H. J.) 685–735 (Chapman & Hall, 1992).
Jensen, B. B. & Burris, R. H. Effect of high pN2 and high pD2 on NH3 production, H2 evolution, and HD formation by nitrogenases. Biochemistry 24, 1141–1147 (1985).
Li, J. L. & Burris, R. H. Influence of pN2 and pD2 on HD formation by various nitrogenases. Biochemistry 22, 4472–4480 (1983).
Burgess, B. K., Wherland, S., Newton, W. E. & Stiefel, E. I. Nitrogenase reactivity: insight into the nitrogen-fixing process through hydrogen-inhibition and HD-forming reactions. Biochemistry 20, 5140–5146 (1981).
Wherland, S., Burgess, B. K., Stiefel, E. I. & Newton, W. E. Nitrogenase reactivity: effects of component ratio on electron flow and distribution during nitrogen fixation. Biochemistry 20, 5132–5140 (1981).
Chatt, J. In Nitrogen Fixation (eds Stewart, W. D. P. & Gallon, J. R.) 1–18 (Academic Press, 1980).
Yang, Z.-Y. et al. On reversible H2 loss upon N2 binding to FeMo-cofactor of nitrogenase. Proc. Natl Acad. Sci. USA 110, 16327–16332 (2013).
Nielander, A. C. et al. A versatile method for ammonia detection in a range of relevant electrolytes via direct nuclear magnetic resonance techniques. ACS Catal. 9, 5797–5802 (2019).
Sippel, D. et al. A bound reaction intermediate sheds light on the mechanism of nitrogenase. Science 359, 1484–1489 (2018).
Tanifuji, K. et al. Tracing the incorporation of the “ninth sulfur” into the nitrogenase cofactor precursor with selenite and tellurite. Nat. Chem. 13, 1228–1234 (2021).
Jasniewski, A. J. et al. Spectroscopic characterization of an eight-iron nitrogenase cofactor precursor that lacks the “9th sulphur”. Angew. Chem. Int. Ed. Engl. 58, 14703–14707 (2019).
Tanifuji, K. et al. Tracing the “ninth sulphur” of the nitrogenase cofactor via a semi-synthetic approach. Nat. Chem. 10, 568–572 (2018).
Wei, W. J. & Siegbahn, P. E. M. A mechanism for nitrogenase including loss of a sulphide. Chemistry 28, e202103745 (2022).
Vincent, K. A. et al. Instantaneous, stoichiometric generation of powerfully reducing states of protein active sites using Eu(II) and polyaminocarboxylate ligands. Chem. Commun. 20, 2590–2591 (2003).
Ribbe, M. W., Hu, Y., Guo, M., Schmid, B. & Burgess, B. K. The FeMoco-deficient MoFe protein produced by a nifH deletion strain of Azotobacter vinelandii shows unusual P-cluster features. J. Biol. Chem. 277, 23469–23476 (2002).
Burgess, B. K., Jacobs, D. B. & Stiefel, E. I. Large-scale purification of high activity Azotobacter vinelandii nitrogenase. Biochim. Biophys. Acta 614, 196–209 (1980).
Lee, C. C., Ribbe, M. W. & Hu, Y. Purification of nitrogenase proteins. Methods Mol. Biol. 1876, 111–124 (2019).
Gavini, N. & Burgess, B. K. FeMo cofactor synthesis by a nifH mutant with altered MgATP reactivity. J. Biol. Chem. 267, 21179–21186 (1992).
Corbin, J. L. Liquid chromatographic-fluorescence determination of ammonia from nitrogenase reactions: a 2-min assay. Appl. Environ. Microbiol. 47, 1027–1030 (1984).
Lee, C. C., Hu, Y. & Ribbe, M. W. Tracing the hydrogen source of hydrocarbons formed by vanadium nitrogenase. Angew. Chem. Int. Ed. Engl. 50, 5545–5547 (2011).
Lee, C. C., Hu, Y. & Ribbe, M. W. Reduction and condensation of aldehydes by the isolated cofactor of nitrogenase. ACS Cent. Sci. 4, 1430–1435 (2018).
Fulmer, G. R. et al. NMR chemical shifts of trace impurities: common laboratory solvents, organics, and gases in deuterated solvents relevant to the organometallic chemist. Organometallics 29, 2176–2179 (2010).
Wiig, J. A., Hu, Y. & Ribbe, M. W. NifEN-B complex of Azotobacter vinelandii is fully functional in nitrogenase FeMo cofactor assembly. Proc. Natl Acad. Sci. USA 108, 8623–8627 (2011).
Schmid, B. et al. Structure of a cofactor-deficient nitrogenase MoFe protein. Science 296, 352–356 (2002).
Acknowledgements
This work was supported by NIH-NIGMS grant no. GM141046 (to Y.H. and M.W.R.). We thank P. R. Dennison, Director of the NMR Facility at UC Irvine, for his kind help with frequency-selective NMR analysis of our samples. We also thank the staff at the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Beamline 12-2 (X-ray diffraction) and Beamlines 7-3 and 9-3 (X-ray absorption spectroscopy) for technical support with data collection. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under contract no. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the US Department of Energy, Office of Biological and Environmental Research and by the National Institutes of Health, National Institutes of General Medical Sciences (no. P30GM133894).
Author information
Authors and Affiliations
Contributions
C.C.L., W.K., A.J.J., M.T.S., K.T., M.W.R. and Y.H. designed experiments. C.C.L., W.K., A.J.J., M.T.S., K.T., M.W.R. and Y.H. analysed data. C.C.L., W.K., A.J.J., M.T.S. and K.T. performed experiments. M.W.R. and Y.H. wrote the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Catalysis thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, Figs. 1–9, Tables 1–9 and references.
Source data
Source Data Fig. 1
Source Data Fig. 1a–c.
Source Data Fig. 2
Source Data Fig. 2a–o.
Source Data Fig. 3
Source Data Fig. 3a–c.
Source Data Fig. 5
Source Data Fig. 5a–d.
Source Data Fig. 6
Source Data Fig. 6a–g.
Source Data Fig. 7
Source Data Fig. 7a–d.
Rights and permissions
About this article
Cite this article
Lee, C.C., Kang, W., Jasniewski, A.J. et al. Evidence of substrate binding and product release via belt-sulfur mobilization of the nitrogenase cofactor. Nat Catal 5, 443–454 (2022). https://doi.org/10.1038/s41929-022-00782-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41929-022-00782-7
This article is cited by
-
Structural insights into the iron nitrogenase complex
Nature Structural & Molecular Biology (2024)
-
Iron-only Fe-nitrogenase underscores common catalytic principles in biological nitrogen fixation
Nature Catalysis (2023)
-
Electrochemical carbon–carbon coupling with enhanced activity and racemate stereoselectivity by microenvironment regulation
Nature Communications (2023)
-
Connecting the geometric and electronic structures of the nitrogenase iron–molybdenum cofactor through site-selective 57Fe labelling
Nature Chemistry (2023)
-
Nitrogenase loosens its belt to fix dinitrogen
Nature Catalysis (2022)