Abstract
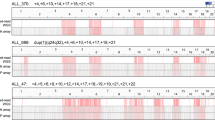
Structural analysis of small supernumerary marker chromosomes (sSMCs) has revealed that many have complex structures. Structural analysis of sSMCs by whole genome sequencing using short-read sequencers is challenging however because most present with a low level of mosaicism and consist of a small region of the involved chromosome. In this present study, we applied adaptive sampling using nanopore long-read sequencing technology to enrich the target region and thereby attempted to determine the structure of two sSMCs with complex structural rearrangements previously revealed by cytogenetic microarray. In adaptive sampling, simple specification of the target region in the FASTA file enables to identify whether or not the sequencing DNA is included in the target, thus promoting efficient long-read sequencing. To evaluate the target enrichment efficiency, we performed conventional pair-end short-read sequencing in parallel. Sequencing with adaptive sampling achieved a target enrichment at about a 11.0- to 11.5-fold higher coverage rate than conventional pair-end sequencing. This enabled us to quickly identify all breakpoint junctions and determine the exact sSMC structure as a ring chromosome. In addition to the microhomology and microinsertion at the junctions, we identified inverted repeat structure in both sSMCs, suggesting the common generation mechanism involving replication impairment. Adaptive sampling is thus an easy and beneficial method of determining the structures of complex chromosomal rearrangements.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Mahmoud M, Gobet N, Cruz-Dávalos DI, Mounier N, Dessimoz C, Sedlazeck FJ. Structural variant calling: the long and the short of it. Genome Biol. 2019;20:246.
Ho SS, Urban AE, Mills RE. Structural variation in the sequencing era. Nat Rev Genet. 2020;21:171–89.
Zepeda-Mendoza CJ, Morton CC. The Iceberg under Water: Unexplored Complexity of Chromoanagenesis in Congenital Disorders. Am J Hum Genet. 2019;104:565–77.
Koltsova AS, Pendina AA, Efimova OA, Chiryaeva OG, Kuznetzova TV, Baranov VS. On the Complexity of Mechanisms and Consequences of Chromothripsis: An Update. Front Genet. 2019;10:393.
Liehr T, Claussen U, Starke H. Small supernumerary marker chromosomes (sSMC) in humans. Cytogenet Genome Res. 2004;107:55–67.
Malvestiti F, De Toffol S, Grimi B, Chinetti S, Marcato L, Agrati C, et al. De novo small supernumerary marker chromosomes detected on 143,000 consecutive prenatal diagnoses: chromosomal distribution, frequencies, and characterization combining molecular cytogenetics approaches. Prenat Diagn. 2014;34:460–8.
Liehr T, Weise A. Frequency of small supernumerary marker chromosomes in prenatal, newborn, developmentally retarded and infertility diagnostics. Int J Mol Med. 2007;19:719–31.
Liehr T. Characterization of prenatally assessed de novo small supernumerary marker chromosomes by molecular cytogenetics. Methods Mol Biol. 2008;444:27–38.
Kotzot D. Complex and segmental uniparental disomy (UPD): review and lessons from rare chromosomal complements. J Med Genet. 2001;38:497–507.
Liehr T, Ewers E, Hamid AB, Kosyakova N, Voigt M, Weise A, et al. Small supernumerary marker chromosomes and uniparental disomy have a story to tell. J Histochem Cytochem. 2011;59:842–8.
Kurtas NE, Xumerle L, Leonardelli L, Delledonne M, Brusco A, Chrzanowska K, et al. Small supernumerary marker chromosomes: a legacy of trisomy rescue? Hum Mutat. 2019;40:193–200.
Matsubara K, Yanagida K, Nagai T, Kagami M, Fukami M. De novo small supernumerary marker chromosomes arising from partial trisomy rescue. Front Genet. 2020;11:132.
Jafari-Ghahfarokhi H, Moradi-Chaleshtori M, Liehr T, Hashemzadeh-Chaleshtori M, Teimori H, Ghasemi-Dehkordi P. Small supernumerary marker chromosomes and their correlation with specific syndromes. Adv Biomed Res. 2015;4:140.
O’Connell J, Schulz-Trieglaff O, Carlson E, Hims MM, Gormley NA, Cox AJ. NxTrim: optimized trimming of Illumina mate pair reads. Bioinformatics. 2015;31:2035–7.
Li H, Durbin R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics. 2010;26:589–95.
Fan X, Abbott TE, Larson D, Chen K. BreakDancer: Identification of Genomic Structural Variation from Paired-End Read Mapping. Curr Protoc Bioinforma. 2014;45(1-11):15.6.
Thorvaldsdóttir H, Robinson JT, Mesirov JP. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform. 2013;14:178–92.
Li H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics. 2018;34:3094–100.
Jiang T, Liu Y, Jiang Y, Li J, Gao Y, Cui Z, et al. Long-read-based human genomic structural variation detection with cuteSV. Genome Biol. 2020;21:189.
Kato T, Inagaki H, Miyai S, Suzuki F, Naru Y, Shinkai Y, et al. The involvement of U-type dicentric chromosomes in the formation of terminal deletions with or without adjacent inverted duplications. Hum Genet. 2020;139:1417–27.
Nurk S, Koren S, Rhie A, Rautiainen M, Bzikadze AV, Mikheenko A, et al. The complete sequence of a human genome. bioRxiv. 2021:2021.05.26.445798.
Zhou L, Zheng Z, Wu L, Xu C, Wu H, Xu X, et al. Molecular delineation of small supernumerary marker chromosomes using a single nucleotide polymorphism array. Mol Cytogenet. 2020;13:19.
Kato T, Ouchi Y, Inagaki H, Makita Y, Mizuno S, Kajita M, et al. Genomic characterization of chromosomal insertions: insights into the mechanisms underlying chromothripsis. Cytogenet Genome Res. 2017;153:1–9.
Bonaglia MC, Kurtas NE, Errichiello E, Bertuzzo S, Beri S, Mehrjouy MM, et al. De novo unbalanced translocations have a complex history/aetiology. Hum Genet. 2018;137:817–29.
Hermetz KE, Newman S, Conneely KN, Martin CL, Ballif BC, Shaffer LG, et al. Large inverted duplications in the human genome form via a fold-back mechanism. PLoS Genet. 2014;10:e1004139.
Crasta K, Ganem NJ, Dagher R, Lantermann AB, Ivanova EV, Pan Y, et al. DNA breaks and chromosome pulverization from errors in mitosis. Nature. 2012;482:53–8.
Zhang CZ, Spektor A, Cornils H, Francis JM, Jackson EK, Liu S, et al. Chromothripsis from DNA damage in micronuclei. Nature. 2015;522:179–84.
Acknowledgements
We thank the patients and their families for agreeing to participate in this study.
Funding
This study was supported by a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science, and Technology (21K06283 to TK, 15H04710 and 24390085 to HK) and from the Ministry of Health, Welfare and Labor (16ek0109067h0003 to HK), Japan.
Author information
Authors and Affiliations
Contributions
TM—Participated in the study design, conducted the molecular experiments, and drafted the manuscript. TK—Participated in the study design, conducted the molecular experiments, and drafted the manuscript. TO—Participated in the study design, conducted the molecular experiments. TS—Participated in the study design, conducted the molecular experiments. SM (S.Miyai)—Participated in the study design, conducted the molecular experiments. HI—Participated in the study design, conducted the molecular experiments. ES—Participated in the study design, conducted the molecular experiments. YM—Participated in the study design, conducted the molecular experiments. SM (S.Mizuno)—Participated in the study design, conducted the molecular experiments. HK—Coordination and conception of the study, critical revision of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The study was approved by the Ethics Committee for Human Genome Studies at Fujita Health University. Written informed consent to participate was obtained from the study patients and their families.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Mariya, T., Kato, T., Sugimoto, T. et al. Target enrichment long-read sequencing with adaptive sampling can determine the structure of the small supernumerary marker chromosomes. J Hum Genet 67, 363–368 (2022). https://doi.org/10.1038/s10038-021-01004-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-021-01004-x
This article is cited by
-
Assessing the efficacy of target adaptive sampling long-read sequencing through hereditary cancer patient genomes
npj Genomic Medicine (2024)
-
DeepSelectNet: deep neural network based selective sequencing for oxford nanopore sequencing
BMC Bioinformatics (2023)
-
Best Practices in Microbial Experimental Evolution: Using Reporters and Long-Read Sequencing to Identify Copy Number Variation in Experimental Evolution
Journal of Molecular Evolution (2023)