Abstract
The efficacy of poly(ADP)-ribose polymerase 1 inhibition (PARPi) in BRCA1-deficient cells depends on 53BP1 and shieldin, which have been proposed to limit single-stranded DNA at double-strand breaks (DSBs) by blocking resection and/or through CST–Polα–primase-mediated fill-in. We show that primase (like 53BP1–shieldin and CST–Polα) promotes radial chromosome formation in PARPi-treated BRCA1-deficient cells and demonstrate shieldin–CST–Polα–primase-dependent incorporation of BrdU at DSBs. In the absence of 53BP1 or shieldin, radial formation in BRCA1-deficient cells was restored by the tethering of CST near DSBs, arguing that in this context, shieldin acts primarily by recruiting CST. Furthermore, a SHLD1 mutant defective in CST binding (SHLD1Δ) was non-functional in BRCA1-deficient cells and its function was restored after reconnecting SHLD1Δ to CST. Interestingly, at dysfunctional telomeres and at DNA breaks in class switch recombination where CST has been implicated, SHLD1Δ was fully functional, perhaps because these DNA ends carry CST recognition sites that afford SHLD1-independent binding of CST. These data establish that in BRCA1-deficient cells, CST–Polα–primase is the major effector of shieldin-dependent DSB processing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All data that were generated and/or analysed in this study are included in the published paper and its Supplementary Information. Conservation symbols according to UniProt can be found at https://www.uniprot.org/. Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding author upon reasonable request.
References
Mirman, Z. & de Lange, T. 53BP1: a DSB escort. Genes Dev. 34, 7–23 (2020).
Pannunzio, N. R., Watanabe, G. & Lieber, M. R. Nonhomologous DNA end-joining for repair of DNA double-strand breaks. J. Biol. Chem. 293, 10512–10523 (2018).
Wright, W. D., Shah, S. S. & Heyer, W. D. Homologous recombination and the repair of DNA double-strand breaks. J. Biol. Chem. 293, 10524–10535 (2018).
Hustedt, N. & Durocher, D. The control of DNA repair by the cell cycle. Nat. Cell Biol. 19, 1–9 (2016).
Bhattacharyya, A., Ear, U. S., Koller, B. H., Weichselbaum, R. R. & Bishop, D. K. The breast cancer susceptibility gene BRCA1 is required for subnuclear assembly of Rad51 and survival following treatment with the DNA cross-linking agent cisplatin. J. Biol. Chem. 275, 23899–23903 (2000).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Roy, R., Chun, J. & Powell, S. N. BRCA1 and BRCA2: different roles in a common pathway of genome protection. Nat. Rev. Cancer 12, 68–78 (2011).
Bouwman, P. et al. 53BP1 loss rescues BRCA1 deficiency and is associated with triple-negative and BRCA-mutated breast cancers. Nat. Struct. Mol. Biol. 17, 688–695 (2010).
Bunting, S. F. et al. 53BP1 inhibits homologous recombination in Brca1-deficient cells by blocking resection of DNA breaks. Cell 141, 243–254 (2010).
Setiaputra, D. & Durocher, D. Shieldin—the protector of DNA ends. EMBO Rep. 20, e47560 (2019).
Zimmermann, M., Lottersberger, F., Buonomo, S. B., Sfeir, A. & de Lange, T. 53BP1 regulates DSB repair using Rif1 to control 5′ end resection. Science 339, 700–704 (2013).
Escribano-Díaz, C. et al. A cell cycle-dependent regulatory circuit composed of 53BP1–RIF1 and BRCA1–CtIP controls DNA repair pathway choice. Mol. Cell 49, 872–883 (2013).
Feng, L., Fong, K. W., Wang, J., Wang, W. & Chen, J. RIF1 counteracts BRCA1-mediated end resection during DNA repair. J. Biol. Chem. 288, 11135–11143 (2013).
Chapman, J. R. et al. RIF1 is essential for 53BP1-dependent nonhomologous end joining and suppression of DNA double-strand break resection. Mol. Cell 49, 858–871 (2013).
Di Virgilio, M. et al. Rif1 prevents resection of DNA breaks and promotes immunoglobulin class switching. Science 339, 711–715 (2013).
Boersma, V. et al. MAD2L2 controls DNA repair at telomeres and DNA breaks by inhibiting 5′ end resection. Nature 521, 537–540 (2015).
Xu, G. et al. REV7 counteracts DNA double-strand break resection and affects PARP inhibition. Nature 521, 541–544 (2015).
Ghezraoui, H. et al. 53BP1 cooperation with the REV7–shieldin complex underpins DNA structure-specific NHEJ. Nature 560, 122–127 (2018).
Mirman, Z. et al. 53BP1–RIF1–shieldin counteracts DSB resection through CST- and Polα-dependent fill-in. Nature 560, 112–116 (2018).
Gupta, R. et al. DNA repair network analysis reveals Shieldin as a key regulator of NHEJ and PARP inhibitor sensitivity. Cell 173, 972–988.e23 (2018).
Dev, H. et al. Shieldin complex promotes DNA end-joining and counters homologous recombination in BRCA1-null cells. Nat. Cell Biol. 20, 954–965 (2018).
Noordermeer, S. M. et al. The shieldin complex mediates 53BP1-dependent DNA repair. Nature 560, 117–121 (2018).
Gao, S. et al. An OB-fold complex controls the repair pathways for DNA double-strand breaks. Nat. Commun. 9, 3925 (2018).
Findlay, S. et al. SHLD2/FAM35A co-operates with REV7 to coordinate DNA double-strand break repair pathway choice. EMBO J. 37, e100158 (2018).
Natsume, T., Kiyomitsu, T., Saga, Y. & Kanemaki, M. T. Rapid protein depletion in human cells by auxin-inducible degron tagging with short homology donors. Cell Rep. 15, 210–218 (2016).
Tang, J. et al. Acetylation limits 53BP1 association with damaged chromatin to promote homologous recombination. Nat. Struct. Mol. Biol. 20, 317–325 (2013).
Han, T. et al. The antitumor toxin CD437 is a direct inhibitor of DNA polymerase α. Nat. Chem. Biol. 12, 511–515 (2016).
Holzer, S. et al. Structural basis for inhibition of human primase by arabinofuranosyl nucleoside analogues fludarabine and vidarabine. ACS Chem. Biol. 14, 1904–1912 (2019).
Nakamura, A. et al. Techniques for gamma-H2AX detection. Methods Enzymol. 409, 236–250 (2006).
Cesare, A. J. et al. Spontaneous occurrence of telomeric DNA damage response in the absence of chromosome fusions. Nat. Struct. Mol. Biol. 16, 1244–1251 (2009).
Cesare, A. J., Heaphy, C. M. & O’Sullivan, R. J. Visualization of telomere integrity and function in vitro and in vivo using immunofluorescence techniques. Curr. Protoc. Cytom. 73, 12.40.1–12.40.31 (2015).
Celli, G. B. & de Lange, T. DNA processing is not required for ATM-mediated telomere damage response after TRF2 deletion. Nat. Cell Biol. 7, 712–718 (2005).
Erhart, D. et al. Chemical development of intracellular protein heterodimerizers. Chem. Biol. 20, 549–557 (2013).
Franco, S. et al. H2AX prevents DNA breaks from progressing to chromosome breaks and translocations. Mol. Cell 21, 201–214 (2006).
Barazas, M. et al. The CST complex mediates end protection at double-strand breaks and promotes PARP inhibitor sensitivity in BRCA1-deficient Cells. Cell Rep. 23, 2107–2118 (2018).
de Lange, T. Shelterin-mediated telomere protection. Annu. Rev. Genet. 52, 223–247 (2018).
Lottersberger, F., Bothmer, A., Robbiani, D. F., Nussenzweig, M. C. & de Lange, T. Role of 53BP1 oligomerization in regulating double-strand break repair. Proc. Natl Acad. Sci. USA 110, 2146–2151 (2013).
Hom, R. A. & Wuttke, D. S. Human CST prefers G-rich but not necessarily telomeric sequences. Biochemistry 56, 4210–4218 (2017).
Chaudhuri, J. & Alt, F. W. Class-switch recombination: interplay of transcription, DNA deamination and DNA repair. Nat. Rev. Immunol. 4, 541–552 (2004).
Anderson, B. H. et al. Mutations in CTC1, encoding conserved telomere maintenance component 1, cause Coats plus. Nat. Genet 44, 338–342 (2012).
Ochs, F. et al. 53BP1 fosters fidelity of homology-directed DNA repair. Nat. Struct. Mol. Biol. 23, 714–721 (2016).
Schimmel, J., Muñoz-Subirana, N., Kool, H., van Schendel, R. & Tijsterman, M. Small tandem DNA duplications result from CST-guided Pol α-primase action at DNA break termini. Nat. Commun. 12, 4843 (2021).
Paiano, J. et al. Role of 53BP1 in end protection and DNA synthesis at DNA breaks. Genes Dev. 35, 19–20 (2021).
Xu, X. et al. Centrosome amplification and a defective G2-M cell cycle checkpoint induce genetic instability in BRCA1 exon 11 isoform-deficient cells. Mol. Cell 3, 389–395 (1999).
Wu, P., Takai, H. & de Lange, T. Telomeric 3′ overhangs derive from resection by Exo1 and Apollo and fill-in by POT1b-associated CST. Cell 150, 39–52 (2012).
Yang, Z., Maciejowski, J. & de Lange, T. Nuclear envelope rupture is enhanced by loss of p53 or Rb. Mol. Cancer Res. 15, 1579–1586 (2017).
Mirzoeva, O. K. & Petrini, J. H. DNA damage-dependent nuclear dynamics of the Mre11 complex. Mol. Cell Biol. 21, 281–288 (2001).
Cesare, A. J., Hayashi, M. T., Crabbe, L. & Karlseder, J. The telomere deprotection response is functionally distinct from the genomic DNA damage response. Mol. Cell 51, 141–155 (2013).
Acknowledgements
We thank A. Bianchi and F. Cross for help designing the yeast two-hybrid SHLD1 mutagenesis screen. Z. Yang is thanked for generating p53/RB-null RPE1 cells, and H. Takai for designing CRISPR–Cas9 gene targeting reagents. We thank F. Stoof for help with the metaIF protocol, which was adapted for metaPLA. B. Conti and A. Smogorzewska provided vidarabine triphosphate, and R. Greenberg is thanked for sharing U2OS-FOKI–LacI cells. We thank D. Durocher for BRCA1 KO and isogenic control RPE1 cells. Z.M. is supported by a NCI F99 award (1F99CA245720-01). A.K. is funded by an EMBO Long-Term Fellowship (ALTF 542-2020) and MRC grant (MR/R017549/1) to J.R.C. J.R.C. is supported by Cancer Research UK (C52690/A19270) and Lister Institute Fellowships. This work was supported by grants from the NCI (5 R35 CA210036), the Breast Cancer Research Foundation (BCRF-19-036) and the Melanoma Research Alliance (MRA#577521) to T.d.L. T.d.L. is an American Cancer Society Rose Zarucki Trust Research Professor.
Author information
Authors and Affiliations
Contributions
Z.M. performed most of the experiments. N.K.S. designed and engineered the RPE1-PRIM1-AID cell line and performed some 3′ overhang assays. A.K. performed the CSR experiment under the supervision of J.R.C. Z.M. and T.d.L. conceived the study and wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
T.d.L. is a member of the Scientific Advisory Board of Calico Life Sciences. The remaining authors declare no competing interests.
Peer review information
Nature Cell Biology thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Generation and validation of p53/Rb-deficient RPE1 cells with mAID knocked into both PRIM1 loci (related to Fig. 1).
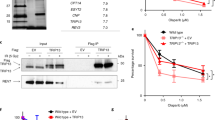
a, Experimental scheme used to tag endogenous PRIM1 with AID-mClover. b, PCR genotyping of mClover-positive clones. Location of primer sets and the expected PCR band sizes are shown in a. c, Immunoblot for mClover-PRIM1 with PRIM1 or anti-GFP antibody. Clone #10, with both copies of PRIM1 tagged with AID-mClover, was selected for the experiments. d, Schematic of HA-OsTIR1 and immunoblot confirming dox-induced HA-OsTIR1 in subclones of PRIM1-AID-mClover clone #10. e, Immunoblot for GFP showing efficient PRIM1 degradation after the indicated IAA (auxin)/dox treatments in two subclones. f, Immunoblot as in e but testing timing and concentration of IAA. g, IF showing loss of EdU incorporation in subclone #6 after auxin/dox-induced PRIM1-AID degradation indicating the expected inhibition of DNA replication. In g, the number of fields of view (n, each represented by a dot) is indicated. In b-g, the experiment has been performed once. Means are indicated with center bars and SDs with error bars.
Extended Data Fig. 2 Primase blocks RAD51 loading in BRCA1-deficient cells (related to Fig. 1).
a, Experimental timeline of auxin-induced degradation of PRIM1 in G2-arrested RPE1 cells. 5 hr after 5 Gy IR, cells were harvested for RAD51 IF. b, c, Quantification of number of RAD51 foci per nucleus for cells with the indicated treatments. This data is summarized in Fig. 1panels e and i. Number of nuclei (n, each represented by a dot) pooled from four independent experiments is indicated. Statistical analyses as in Fig. 1. All means are indicated with center bars and SDs with error bars.
Extended Data Fig. 3 Experimental setup for detection of fill-in synthesis in U2OS-FOKI cells (related to Fig. 2).
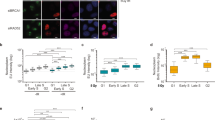
a, Experimental timeline for IF to detect BrdU incorporation at DSBs in G2-arrested U2OS cells. b, Representative IF images of cells as in a visualizing DAPI, 53BP1, mCherry-FOKI, and BrdU. The middle cell displays a global BrdU incorporation pattern, indicative of S phase. Image representative of four independent experiments. Scale bar, 20 μm. c, Experimental timeline and IF protocols for U2OS-FOKI-LacI cells released into G1. d, Representative IF images of HA-STN1 detected in U2OS-FOKI-LacI cells arrested in G2 with RO-3306 or released into G1 (9 μM RO-3306 overnight followed by washout into fresh media for 2 h before induction of FOKI). Nuclear outlines are demarcated by dashed white lines. Scale bars, 5 μm. e, Quantification of HA-STN1 colocalization with FOKI/53BP1 foci in cells as in d. n = three independent experiments. f, Representative IF for Cyclin A in cells as in c. Note the two pairs of small, neighboring, ‘doublet’ cells, which in all cases were Cyclin A-negative. Such cell pairs are likely to represent the two daughter cells from a recent mitosis. Scale bar, 20 μm. g, Quantification of FOKI/BrdU colocalizations in induced, non-S-phase, ‘doublet’ cells as in f after CDKi washout with or without Polα inhibitor. Data from three (CD437) or four (DMSO) independent experiments. Statistical analyses as in Fig. 1. All means are indicated with center bars and SDs with error bars.
Extended Data Fig. 4 MetaPLA for detection of fill-in synthesis in BRCA1-deficient cells (related to Fig. 3).
a, Representative images from metaPLA of BrdU/γH2AX on metaphase spreads in Trf2F/F Lig4−/− MEFs with the indicated treatment that were used to validate the technique. Scale bar, 10 μm. b, Quantification of terminal, chromatin-associated BrdU/γH2AX PLA foci for cells as in a. c, Immunoblot for BRCA1 and REV7 in the indicated cells with or without Cre treatment as in Fig. 3c. Representative of three independent experiments. d, Quantification of chromatin-associated BrdU/γH2AX PLA foci on metaphase spreads in Brca1F/F MEFs of the indicated genotype, with 5 or 1 μM Olaparib. e, BrdU/γH2AX PLA controls quantified as in d. In b, d, and e, n = the number of metaphases scored in each condition as indicated (n) pooled from three independent experiments. Statistical analyses as in Fig. 1. All means are indicated with center bars and SDs with error bars.
Extended Data Fig. 5 Fill-in synthesis occurs late in G2 in BRCA1-deficient cells.
a, Experimental timeline for Polα inhibition in PARPi-treated, BRCA1-deficient MEFs released from G2 into prolonged (180 min) metaphase arrest. b, Quantification of the percent of chromosomes involved in radial structures. Number of metaphases (n, each represented by a dot) pooled from three independent experiments is indicated. c, Representative sample images of nuclei with intact or broken-down nuclear envelopes (NE) as assessed by Lamin A/C IF. Scale bars, 5 μm. d, Top, analysis of NE breakdown timing after release from G2 in cells as in a and c assessed at the indicated time points. Bottom, analysis of RAD51 foci formation in the same population of cells. Data from three independent experiments. For NE breakdown, Lamin A/C integrity was visually assessed in at least 800 cells per condition per experiment. For RAD51 foci formation, 50-136 nuclei were analyzed per condition per experiment. e, Quantification of RAD51-positive nuclei in Brca1F/F MEFs following treatment with Cre and IR. n = three (no Cre) or four (+ Cre) independent experiments. f, Quantification of percent of chromosomes involved in radial structures after the indicated treatment. Number of metaphases (n, each represented by a dot) pooled from four independent experiments is indicated. Statistical analyses as in Fig. 1. All means are indicated with center bars and SDs with error bars.
Extended Data Fig. 6 Bypass of 53BP1/shieldin by artificial tethering of CST (related to Fig. 4).
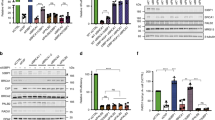
a, Immunoblots for BRCA1, 53BP1, and SHLD1 detecting FHA-SHLD1 in the indicated MEFs expressing FHA-R61Q-SHLD1 (RQ), corresponding to Fig. 4b. b, Immunoblots for BRCA1 and SHLD1 detecting FHA-SHLD1 in the indicated MEFs, corresponding to Fig. 4c. c, Quantification of 53BP1 and GFP-FHA-STN1 foci at DSBs in the indicated MEFs after IR (n = 3 independent experiments, except for the first two columns, where n = 2 independent experiments. d, Representative images of irradiated Brca1F/F Shld2−/− or Brca1F/F 53BP1−/− MEFs as in c. Nuclear outlines are demarcated by dashed white lines. e, Immunoblots for BRCA1, 53BP1, and STN1 detecting endogenous and FHA-STN1 in Brca1F/F Shld2−/− MEFs as in Fig. 4i. f, Representative images of DAPI-stained metaphase spreads from cells as in e, corresponding to Fig. 4i. All scale bars, 10 μm. All panels are representative of three independent experiments. All means are indicated with center bars and SDs with error bars.
Extended Data Fig. 7 Identification of a SHLD1 mutant with impaired CTC1 interaction (related to Fig. 5).
a, Schematic of 53BP1 and its downstream effectors. Interactions (lines) based on previous reports13,18,19,20,21,22,23. Black lines, interactions demonstrated by coimmunoprecipitation; purple lines, interactions demonstrated by yeast two-hybrid19. Asterisk denotes the SHLD1-CTC1 interaction targeted for disruption in the random mutagenesis screen in b. b, Mutants identified in a yeast two-hybrid screen for loss of CTC1 binding executed with randomly mutagenized human SHLD1 ORFs. Left, candidate GAD-SHLD1-URA3 variants (red numbers) grow on permissive (-leucine, -tryptophan, -uracil) media. Expression of full-length SHLD1 variants is ensured by growth on media lacking uracil. Several controls are also shown (black numbers, for example, c1a). Variants that fail to grow on selective (-leucine, -tryptophan, -uracil, -adenine) media are indicated with a green circle (or triangle for deletion). Five sequenced variants were attributable to a single mutation (green shape with magenta border). Mutation L20P, deletion of nearby A23, or the mutations A119V or Q127L severely diminished the SHLD1-CTC1 interaction. Right, sequence alignment of human and mouse SHLD1 with conserved residues highlighted in yellow. SHLD1 sequence variants are represented graphically by shapes above the sequence (see legend at bottom). The experiment has been performed once. c, Multiple sequence alignment of SHLD1. Conservation symbols are according to Uniprot: asterisks, fully conserved; colon, strong similarity; period, weak similarity. The orange outline highlights the conserved LDLP motif deleted in the SHLD1Δ mutant used in this study.
Extended Data Fig. 8 Characterization of SHLD1∆ (related to Fig. 5).
a, Immunoblots showing expression of SNAP-SHLD1 or SNAP-SHLD1∆LDLP (∆) in U2OS cells. b, Quantification of SNAP-SHLD1 localization to IR-induced γH2AX foci in cells as in a. n = 45 nuclei pooled from three independent experiments. c, Immunoprecipitation of myc-SHLD2C (aa 421-904) and immunoblot for SNAP-SHLD1 co-expressed in 293 T cells. d, Representative IF images showing γH2AX co-localizing with HA-STN1 in irrradiated BRCA1/SHLD1 DKO cells complemented with wt SHLD1 or SHLD1∆. Nuclear outlines are demarcated by dashed white lines. Scale bars, 5 μm. Five sample foci are shown for each nucleus. e, Quantification of IR-induced γH2AX foci with HA-STN1 signal in the indicated cells. Red dotted line: the average background level across multiple conditions of random overlaps between γH2AX and HA foci (see Materials and Methods). f, Quantification of γH2AX foci in the indicated RPE1 cells with HA-STN1 as in d and e. Center bar indicates median. g, Quantification of RAD51 foci as in Fig. 5e in parental (wt), BRCA1 KO, or SHLD1 KO RPE1 cells. n = 3-7 independent experiments (as indicated by the number of data points). Ordinary one-way ANOVA was performed with Tukey’s correction for multiple comparisons in g. h, Quantification of the number of RAD51 foci per nucleus for cells as in Fig. 5e with the indicated FLAG-SHLD1 constructs. In e, f, and h, the number of nuclei (n, each represented by a dot) pooled from three independent experiments is indicated. i, Representative images of DAPI-stained metaphase spreads (top; orange arrows denote aberrant radial chromosomes) or RAD51 foci (bottom; nuclear outlines demarcated by dashed white lines) in BRCA1/SHLD1 DKO cells complemented with an empty vector control, wt SHLD1, or SHLD1∆. Scale bars, 5 μm. a and c are representative of two independent experiments.. Statistical analyses as in Fig. 1. All means are indicated with center bars (unless otherwise noted) and SDs with error bars.
Extended Data Fig. 9 SHLD1∆ suppresses overhang length at TPP1-deficient telomeres (related to Fig. 6).
a, Immunoblot showing expression of 53BP1 pathway components and SHLD1 construct expression in Tpp1F/F Shld1+/+ or Shld1−/− clones. b, Immunoblot showing bulk CRISPR KO of BRCA1 and SHLD1 construct expression in cells of the indicated genotype with SHLD1 construct expression. c, Quantitative analysis of telomeric ss overhang intensity in cells as in b using in-gel hybridization to detect the 3’ overhang followed by rehybridization to the denatured DNA in the same gel, allowing the ratio of ss to total TTAGGG signal to be determined. d, Immunoprecipitation of SHLD1 in Tpp1F/F MEFs with the indicated treatments and STN1 immunoblot. Experiments in a-d have been performed once.
Extended Data Fig. 10 SHLD1∆ suppresses overhang length at TRF2-deficient telomeres (related to Fig. 6).
a, Immunoblot showing SHLD1 construct expression and TRF2 deletion by Cre in Trf2F/F Lig4−/− Shld1+/+ or Shld1−/− clones. Asterisks indicate non-specific bands. b, Quantitative analysis of telomeric ss overhang intensity in cells as in a. c, Quantification of overhangs from Trf2F/F Lig4−/− Shld1−/− cells (n = 3 independent experiments. a-c represent data from three independent experiments using two independent clones (circles and triangle). d, Immunoblot showing bulk CRISPR KO of BRCA1 (arrow), phosphorylation of CHK2 after TRF2 deletion, and SHLD1 construct expression. e, Telomeric overhang analysis on cells as in d. Statistical analysis was performed using two-tailed ratio-paired t-test. *, p < 0.05; **, p < 0.01; ns, not significant. Experiments in d, e have been performed once. f, Schematic of mouse IgA Switch region with several five nucleotide repeat sequences and their number of repeats in the region (4.4 kb). All means are indicated with center bars and SDs with error bars.
Supplementary information
Supplementary Information
Supplementary Fig. 1: Gating strategy used in Fig. 6a.
Source data
Source Data Fig. 1
Statistical numerical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 2
Statistical numerical source data.
Source Data Fig. 2
Unprocessed western blots.
Source Data Fig. 3
Statistical numerical source data.
Source Data Fig. 3
Unprocessed western blots.
Source Data Fig. 4
Statistical numerical source data.
Source Data Fig. 4
Unprocessed western blots.
Source Data Fig. 5
Statistical numerical source data.
Source Data Fig. 5
Unprocessed western blots.
Source Data Fig. 6
Statistical numerical source data.
Source Data Fig. 6
Unprocessed western blots.
Source Data Extended Data Fig. 1
Statistical numerical source data.
Source Data Extended Data Fig. 1
Unprocessed western blots.
Source Data Extended Data Fig. 2
Statistical numerical source data.
Source Data Extended Data Fig. 3
Statistical numerical source data.
Source Data Extended Data Fig. 4
Statistical numerical source data.
Source Data Extended Data Fig. 4
Unprocessed Western blots.
Source Data Extended Data Fig. 5
Statistical numerical source data.
Source Data Extended Data Fig. 6
Statistical numerical source data.
Source Data Extended Data Fig. 6
Unprocessed Western blots.
Source Data Extended Data Fig. 8
Statistical numerical source data.
Source Data Extended Data Fig. 8
Unprocessed Western blots.
Source Data Extended Data Fig. 9
Unprocessed western blots.
Source Data Extended Data Fig. 10
Statistical numerical source data.
Source Data Extended Data Fig. 10
Unprocessed western blots.
Rights and permissions
About this article
Cite this article
Mirman, Z., Sasi, N.K., King, A. et al. 53BP1–shieldin-dependent DSB processing in BRCA1-deficient cells requires CST–Polα–primase fill-in synthesis. Nat Cell Biol 24, 51–61 (2022). https://doi.org/10.1038/s41556-021-00812-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00812-9
This article is cited by
-
RTF2 controls replication repriming and ribonucleotide excision at the replisome
Nature Communications (2024)
-
Targeting DNA damage response pathways in cancer
Nature Reviews Cancer (2023)
-
Dynamics of the DYNLL1–MRE11 complex regulate DNA end resection and recruitment of Shieldin to DSBs
Nature Structural & Molecular Biology (2023)
-
SHLD1 is dispensable for 53BP1-dependent V(D)J recombination but critical for productive class switch recombination
Nature Communications (2022)
-
Cryo-EM structure of the human CST–Polα/primase complex in a recruitment state
Nature Structural & Molecular Biology (2022)